 Marquès Lab
Marquès Lab
 Computational Biology and Health Genomics
Computational Biology and Health Genomics
- Group page
- Research lines
- Group members
- Publications
- Datasets
- Software
2026, Affiliated Group Leader, Centre for Genomic Regulation, Barcelona, Spain
2019 - today, Part time Full Professor, Universitat Pompeu Fabra, Barcelona, Spain
2016 - 2020, Director, Institute of Evolutionary Biology (IBE, CSIC-UPF), Barcelona, Spain
2011 - today, ICREA Research Professor
2010 - 2011, Ramon y Cajal Fellow
2010 - today, Principal Investigator, Institute of Evolutionary Biology (IBE, CSIC-UPF), Barcelona, Spain
2007 - 2010, Marie Curie postdoctoral fellow, University of Washington, Seattle, USA
2002 - 2007, PhD Student, Universitat Pompeu Fabra, Barcelona, Spain
Summary
Over the past 15 years, the group has focused on understanding human biology through primate genomics. By comparing the human genome with those of other primates, the lab investigates how evolutionary changes have shaped human-specific traits, disease susceptibility, and brain function. The research integrates evolutionary biology, functional genomics, neuroscience, and conservation science.
In 2023, the group contributed to a Science Special Issue reporting the sequencing of high-quality genomes for 50% of the world’s primate species, an unprecedented international effort that generated transformative resources for evolutionary and biomedical research. These data provide a comparative framework to interpret human genetic variation, revealing how mutations accumulated across primate lineages help predict functional consequences in the human genome. In 2024, the team co-led a Nature study on the evolution of non-coding regions, highlighting the role of regulatory changes in human-specific biology.
The group previously demonstrated that structural variation accumulated differently in the human lineage compared to other great apes (Nature, 2009) and led the first global analysis of great ape genome diversity (Nature, 2013), reconstructing demographic history and genetic differentiation across species. More recently, the lab recovered the oldest ape genetic material, identified introgression into bonobos from an extinct chimpanzee lineage, and developed methods to obtain nuclear DNA from non-invasive samples—advancing both evolutionary research and conservation.
Beyond genome sequence comparisons, the group characterizes regulatory and epigenetic differences between humans and great apes, including DNA methylation, histone modifications, mRNA splicing, and protein isoform diversity. In collaboration with colleagues at Yale, the lab generated the most comprehensive molecular comparison of human and chimpanzee brains to date.
With over 200 peer-reviewed publications and major competitive funding, including ERC and NIH grants, the group bridges zoology and molecular biomedicine, leveraging primate evolution to illuminate the biological foundations of our species.
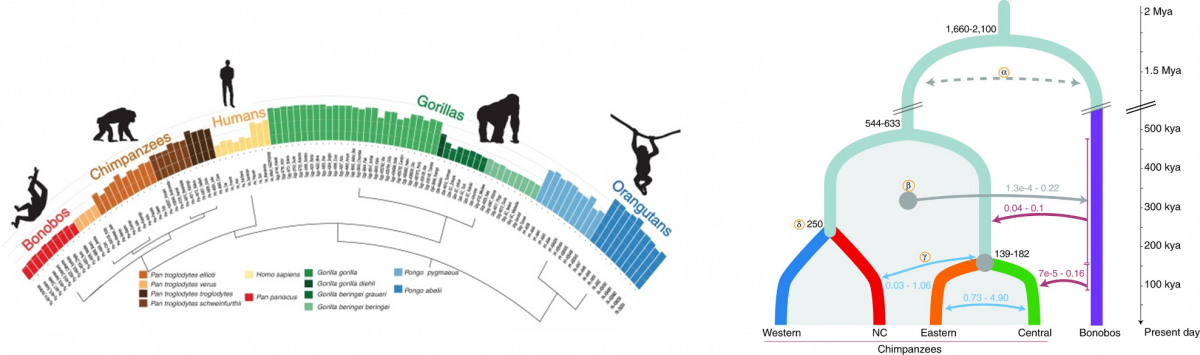
Research Line 1: Primate genetic variation in the context of human evolution and conservation biology.
Characterizing the variation of thousands of human genomes is standard today. However, primates (our closest relatives) are the ideal set of species to study the evolution of these features from both mechanistic and adaptive points of view. In this line of research, we use genomic approaches in primate genomes to understand the impact of variants in the evolution of every species to provide a proper perspective to the differences among species.
Related Publications:
- Prado-Martinez et al. Nature. 2013
- Xue et al. Science. 2015
- deManuel et al. Science 2016
- Kuhlwilm et al. Nature Ecology and Evolution 2019
- Fontsere et al. Cell Genomics 2022
- Kuderna et al. Science 2023
- Gao et al. Science 2023
- Pawar et al. Nature Ecology and Evolution 2023
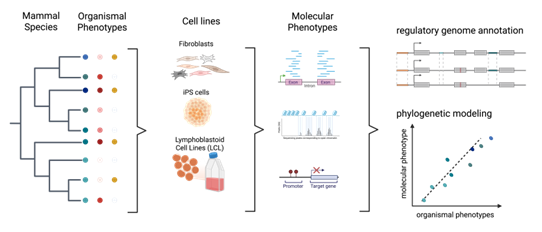
Research Line 2: Comparative gene regulation in humans and primates
DNA methylation is an epigenetic modification involved in regulatory processes such as cell differentiation during development, X-chromosome inactivation, genomic imprinting and susceptibility to complex disease. However, the dynamics of DNA methylation changes between humans and their closest relatives is still poorly understood. In this project, we evaluate methylation patterns in recent human evolution. We identified a significant positive relationship between the rate of coding variation and alterations of methylation at the promoter level.
Related Publications:
- Hernando et al. Plos Genetics 2013
- Hernando et al. NAR 2015
- Hernando et al. Plos Genetics 2015 (Review)
- Garcia et al. Nature Communications 2021
- Ferrandez et al. Genome Research 2022
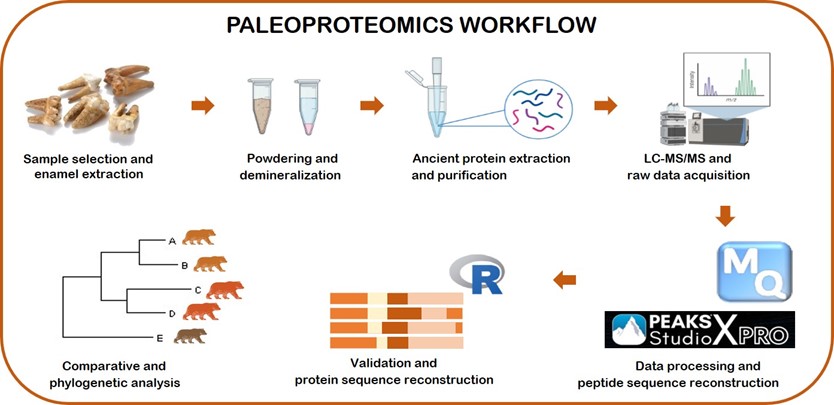
Research Line 3: Paleoproteomics
Paleoproteomics is the scientific field that focuses on the study of ancient proteins preserved in archaeological and paleontological materials. By analyzing protein sequences, paleoproteomics provides unique insights into the evolutionary relationships, dietary practices, health, and biology of ancient organisms, including humans. Our research enhances our understanding of the molecular history of life on Earth, offering a complementary perspective to DNA analysis. Our group is dedicated to advancing paleoproteomic methodologies and applications, aiming to uncover new dimensions of historical biological data particularly in non-human species.
Related Publications:
- Fong-Zazueta, Krueger et al. Bioarxiv 2024. https://www.biorxiv.org/content/10.1101/2024.02.28.580462v1
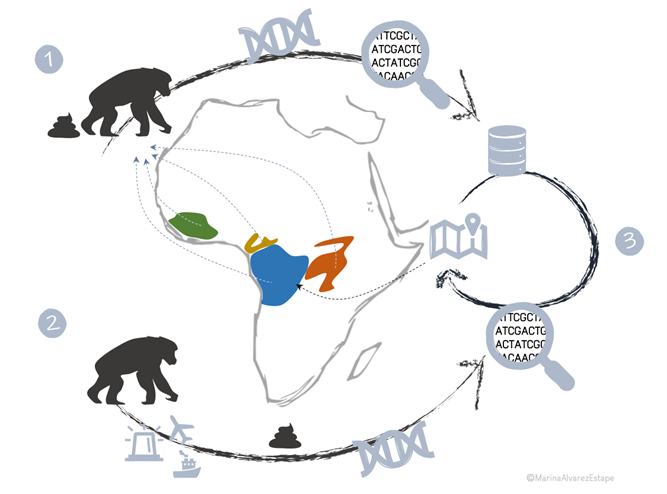
Research Line 4: Primate Conservation: Genetic Monitoring of primate captive population
Thousands of great apes are being illegally transported out of Africa and Asia as part of the orphan ape and bushmeat trades. Detecting the origin of confiscated individuals and bushmeat products is critical in combating wildlife crime. Currently, there is no reliable forensic genetic work for apes that can predict the origin of animals accurately and affordably. Our group has developed a technology that can geolocate chimpanzees within a few hundred km of their origin using genome sequencing from non-invasive samples (Fontsere et al., Cell Genomics, 2022). Analyzing samples from the captive population is a primary goal of this project, and its inferred geolocation data will be cross-referenced with a comprehensive genetic database built from GPS-located samples to generate a comprehensive view on the genetic origins of these samples.
Research Line 5: Biodiversity Monitoring: eDNA
The need for improved biodiversity monitoring tools is evident as we seek to better understand the spatial and temporal distribution of organisms, their behaviors, and their interactions within ecosystems. This understanding is crucial for making informed conservation decisions, shaping strategies for natural resource management, and assessing the impacts of large-scale industrial activities on biodiversity. To address these challenges, innovative technologies such as environmental DNA (eDNA) and bioacoustic monitoring have emerged as powerful tools for biomonitoring. eDNA involves collecting and analyzing genetic material found in the environment, such as water, soil, and air, to identify the species present in a particular area. This method allows for non-invasive and comprehensive monitoring of biodiversity, providing insights into species diversity and distributions. As a part of this line of research, our group has participated in the XPrize Competition: https://www.xprize.org/prizes/rainforest
Research Line 6: Cryozoo: A biobank for biomedicine and conservation
The Cryozoo at the Barcelona Zoological Park is a pioneering biobank dedicated to the conservation and study of animal cell lines, focusing on endangered species. In collaboration with the Pompeu Fabra University, EMBL and Museu Nacional de Historia Natural, the Cryozoo aims to preserve the genetic diversity of species through comprehensive cell line collection and molecular datasets. This initiative supports research in conservation, evolutionary biology, and biomedicine, offering a valuable resource for understanding and protecting biodiversity. The biobank stores well-characterized, karyotyped cell lines, ensuring the availability of crucial molecular data for future research and conservation efforts.



