Functional and evolutionary impact of neural-specific exons and microexons in vertebrates
Functional and evolutionary impact of neural-specific exons and microexons in vertebrates
Current lab members: Patryk Polinski, Tahnee Mackensen, Ludovica Ciampi, Jon Permanyer, Cristina Rodriguez, Luis P. Iñiguez, Marta Miret-Cuesta, Federica Mantica.
Former lab members: Laura López-Blanch, Elisabeth Kita, Antonio Torres-Méndez, Yamile Márquez, Mireia Codina.
Funding: ERC Consolidator Grant, Marie Skłodowska-Curie Actions, EMBO, MICINN, CRG.
Mirroring their unparalleled morphological and cellular complexity, vertebrate brains show the highest levels of regulated alternative splicing known in nature. However, the functions of most of these alternative transcripts, and the evolutionary impact that the increased transcriptional complexity has had on the evolution of the vertebrate brain are still widely unexplored.
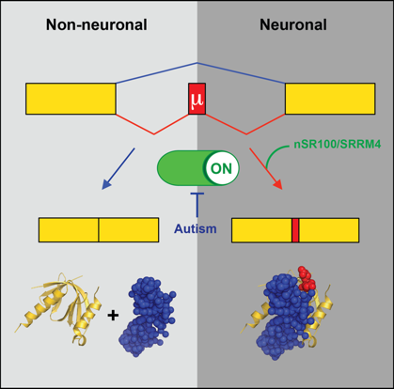
Microexons (Irimia et al, Cell 2014)
In this research line, we are investigating the functions and evolutionary impact of neural-specific alternative splicing in vertebrates. In particular, we focus on microexons, tiny exons between 3-27 nucleotides long that show striking switches during neuronal regulation and unmatched evolutionary conservation across vertebrates. We aim to understand (i) the evolutionary origins of microexons and their span across metazoan nervous systems, (ii) the molecular bases and conservation of their exquisite neuronal regulation, (iii) their impact on protein structures and protein-protein interactions, (iv) their roles during vertebrate embryo development, and (v) their potential causal links with autism spectrum disorder and other human diseases.
In addition, we have recently identified a subset of microexons specifically enriched in photoreceptors, which we termed RetMICs. We are therefore interested in understanding their functions, evolution and relation with microexons in other sensory neurons.

Protein microsurgery by microexons (Alexander Weiss)
To achieve these goals, we apply a combination of bioinformatics and high-throughput transcriptomic analyses in multiple vertebrate and non-vertebrate species, experimental manipulation in model organisms (zebrafish, mouse and fruitfly) using CRISPR-Cas9 technology for genome editing, and systems-level analysis and modelling of proteins structures and interaction networks.
Related publications:
- Ciampi, L., Mantica, F., Lopez-Blanch, L., Rodríguez-Marin, C., Cianferoni, D., Zang, J., Permanyer, J., Jiménez-Delgado, S., Bonnal, S., Miravet-Verde, S., Ruprecht, V., Neuhauss, S.C.F., Banfi, S., Carrella, S., Serrano, L.†, Head, S.A.†, Irimia, M.† (2022). Specialization of the photoreceptor transcriptome by Srrm3-dependent microexons is required for outer segment maintenance and vision. PNAS, 119:e2117090119.
- Torres-Méndez, A.†, Pop, S., Bonnal, S., Almudi, I., Roberts, R.J.V., Paolantoni, C., Alcaina, A., Avola, A., Martín-Anduaga, A., Haussmann, I.U., Morin, V., Casares, F., Soller, M., Kadener, S., Roignant, J.-Y., Prieto-Godino, L.†, Irimia, M.† (2022). Parallel evolution of a splicing program controlling neuronal excitability in flies and mammals. Sci Adv, 8:eabk0445.
- Elorza, A., Márquez, Y., Cabrera, J.R., Sánchez-Trincado, J.L., Santos-Galindo, M., Hernández, I.H., Picó, S., Díaz-Hernández, J.I., García-Escudero, R., Irimia, M.†, Lucas J.J.† (2021). Huntington’s disease-specific mis-splicing unveils key effector genes and altered splicing factors. Brain, 144:2009-23.
- Head, S.A., Hernandez-Alias, X., Yang, J.S., Ciampi, L., Beltran-Sastre, V., Torres-Méndez, A., Irimia, M., Schaefer, M.H., Serrano, L. (2021). Silencing of SRRM4 suppresses microexon inclusion and promotes tumor growth across cancers. PLoS Biol, 19(2):e3001138.
- Gonatopoulos-Pournatzis, T., Niibori, R., Salter, E.W., Weatheritt, R.J., Tsang, B., Farhangmehr, S., Liang, X., Braunschweig, U., Roth, J., Zhang, S., Henderson, T., Sharma, E., Quesnel-Vallières, M., Permanyer, J., Maier, S., Georgiou, J., Irimia, M., Sonenberg, N., Forman-Kay, J.D., Gingras, A.C., Collingridge, G.L., Woodin, M.A., Cordes, S.P., Blencowe, B.J. (2020). Autism-Misregulated eIF4G Microexons Control Synaptic Translation and Higher Order Cognitive Functions. Mol Cell, 77: 1176-1192.
- Torres-Méndez, A., Bonnal, S., Marquez, Y., Roth, J., Iglesias, M., Permanyer, J., Almudí, I., O’Hanlon, D., Guitart, T., Soller, M., Gingras, A.-C., Gebauer, F., Rentzsch, F., Blencowe, B.J.B., Valcárcel, J., Irimia, M.† (2019). A novel protein domain in an ancestral splicing factor drove the evolution of neural microexons. Nature Ecol Evol, 3:691-701.
- Blazquez, L., Emmett, W., Faraway, R., Pineda, J.M.B., Bajew, S., Gohr, A., Haberman, N., Sibley, C.R., Bradley, R.K., Irimia, M., Ule, J. (2018). Exon Junction Complex Shapes the Transcriptome by Repressing Recursive Splicing. Mol Cell, 72:496-509.
- Fernandez, J.P., Moreno-Mateos, M.A., Gohr, A., Miao, L., Chan, S.H., Irimia, M.†, Giraldez, A.J.† (2018). RES complex is associated with intron definition and required for zebrafish early embryogenesis. PLoS Genet, 14:e1007473.
- Parras, A., Anta, H., Santos-Galindo, M., Swarup, V., Elorza, A., Nieto-Gonzalez, J.L., Picó, S., Hernández, I.H., Díaz-Hernández, J.I., Belloc, E., Rodolosse, A., Parikshak, N.N., Peñagarikano, O., Fernández-Chacón, R., Irimia, M., Navarro, P., Geschwind, D.H., Méndez, R., Lucas, J.J. (2018). Autism-like phenotype and risk gene-RNA deadenylation by CPEB4 mis-splicing. Nature, 560:441-446.
- Irimia, M., Weatheritt, R.J., Ellis, J., Parikshak, N.N., Gonatopoulos-Pournatzis, T., Babor, M., Quesnel-Vallières, M., Tapial, J., Raj, B., O’Hanlon, D., Barrios-Rodiles, M., Sternberg, M.J.E., Cordes, S.P., Roth, F.P., Wrana, J.L., Geschwind, D.H., Blencowe, B.B. (2014). A highly conserved program of neuronal microexons is misregulated in autistic brains. Cell, 159:1511-23.