 Sdelci Lab
Sdelci Lab
 Genome Biology
Genome Biology
- Group page
- Research lines
- Group members
- Publications
2005 B.Sc. in Biotechnology, University of Florence, Florence (Italy)
2007 M.Sc. in Medical Biotechnology, University of Florence, Florence (Italy)
2007-2009 Internship at the Department of Pathology and Oncology, University of Florence (Italy)
2012 PhD in Biomedicine, IRB (Institute for Research in Biomedicine), Barcelona
2013-2016 JDRF Postdoctoral fellow at CeMM (Research Center for Molecular Medicine of the Austrian Academy of Sciences), Vienna
2016-2018 Senior postdoctoral researcher at CeMM (Research Center for Molecular Medicine of the Austrian Academy of Sciences), Vienna
2017 M.Sc. in Bioethics, University Ramon Llull, Barcelona (Spain)
January 2019 Group Leader at the Centre for Genomic Regulation (CRG), Barcelona (Spain)
News
Sara Sdelci receives the VI FERO-MANGO Award in Breast Cancer (11/06/2024)
Sara Sdelci, Group Leader at the Centre for Genomic Regulation, has received the VI FERO-MANGO Breast Cancer Project from the FERO Foundation, a prize worth 80,000 euros.
‘Double strike’ strategy slows growth of drug-resistant breast cancer (08/11/2023)
Researchers at the Centre for Genomic Regulation and Vall d’Hebron Institute of Oncology have shown that the simultaneous inhibition of two different proteins may represent a new strategy for tackling triple-negative breast cancer, the most aggressive and drug-resistant form of breast cancer.
DNA damage repaired by antioxidant enzymes (01/06/2023)
In crisis, the nucleus calls antioxidant enzymes to the rescue. The nucleus being metabolically active is a profound paradigm shift with implications for cancer research.
Sara Sdelci awarded the Leonardo Grant from the BBVA Foundation (06/10/2022)
The grant will help determine whether the location of the IMPDH2 enzyme in chromatin (the natural form of DNA inside the cell nucleus) has a metabolic vulnerability in triple negative breast cancer.
CRG scientists receive €5m to research cancer, ageing and evolution (03/09/2019)
Sara Sdelci, a Group Leader running a lab in the CRG’s gene regulation, stem cells and cancer research programme, is one of the recipients. Her project EPICAMENTE will explore the roles of enzymes in cancer proliferation.
Summary
The central role of metabolic rewiring during cancer progression is undeniable, but its direct impact on chromatin functions has been poorly investigated.
We recently described that enzymes of central metabolism can localize on chromatin in cancer cells. However, their chromatin-associated roles remain mostly elusive. Therefore, the scope of our research is to dissect the role of enzymatic activities on chromatin, with a particular focus on cancer. To do so, we apply chemical biology and functional genomics approaches to unbiased chromatin reporters (REDS) for selected metabolic enzymes. We aim to identify the molecular networks defining the direct interplay between chromatin and cancer metabolism. Our research focuses on a brand-new area of cancer biology and has the potential to uncover novel chromatin-associated metabolic vulnerabilities. The breakthrough aspect of our investigation has been recognized by the European Research Council which, in 2019, awarded us with one of the prestigious ERC starting grant (https://erc.europa.eu/projects-figures/erc-funded-projects/results?search_api_views_fulltext=epicamente).
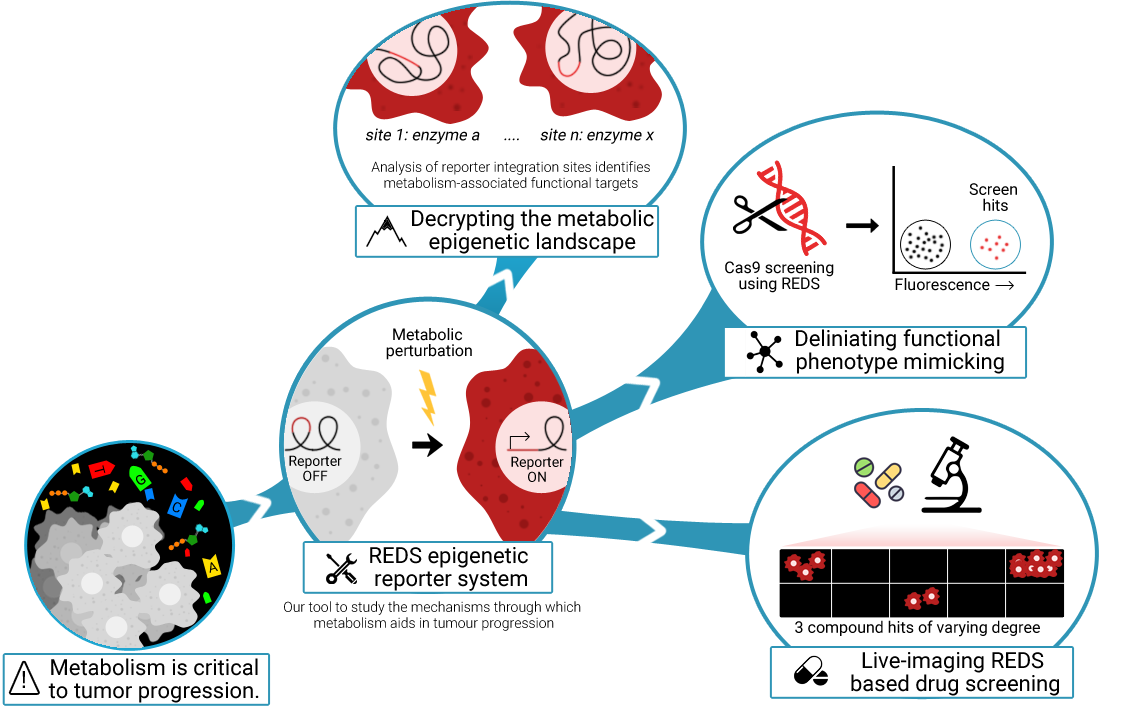
Credit: Rito Ghose
Funding acknowledgements
The project "Decoding chromatin-metabolism in breast cancer" (PID2019-110598GA-I00 / AEI / 10.13039/501100011033) has received funding from the National Research Agency (Agencia Estatal de Investigación, AEI), from the Spanish Ministry of Science and Innovation.





